Heteroduplex
This article needs additional citations for verification. (April 2023) |
A heteroduplex is a double-stranded (duplex) molecule of nucleic acid originated through the genetic recombination of single complementary strands derived from different sources, such as from different homologous chromosomes or even from different organisms.
One such example is the heteroduplex DNA strand formed in hybridization processes, usually for biochemistry-based phylogenetic analyses. Another is the heteroduplexes formed when non-natural analogs of nucleic acids are used to bind with nucleic acids; these heteroduplexes result from performing antisense techniques using single-stranded peptide nucleic acid, 2'-O-methyl phosphorothioate or Morpholino oligos to bind with RNA.
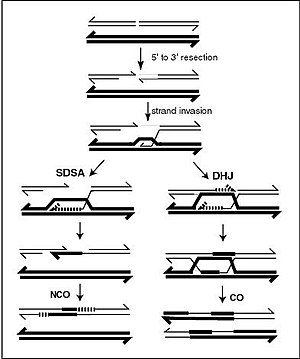
Meiotic recombination can be initiated by a double-strand break (DSB) in DNA. The 5’ ends of the break are degraded, leaving long 3’ overhangs of several hundred nucleotides (see Figure). One of these 3’ single stranded DNA segments then invades a homologous sequence on the homologous chromosome, forming an intermediate which can be repaired through different pathways resulting either in crossovers (CO) or noncrossovers (NCO) as illustrated in the Figure. By one pathway, a structure called a double Holliday junction (DHJ) is formed, leading to the exchange of DNA strands. By the other pathway, referred to as Synthesis-dependent strand annealing (SDSA), there is information exchange but not physical exchange. At various steps of these recombination processes, heteroduplex DNA (double-stranded DNA consisting of single strands from each of the two homologous chromosomes which may or may not be perfectly complementary) is formed. During meiosis non-crossover recombinants occur frequently and these appear to arise mainly by the SDSA pathway.[1][2] Non-crossover recombination events occurring during meiosis likely reflect instances of repair of DNA double-strand damages or other types of DNA damages. When mismatches occur in heteroduplex DNA, the sequence of one strand can be repaired to bind the other strand with perfect complementarity.
During mitosis, the major homologous recombination pathway for repairing DNA double-strand breaks appears to be the SDSA pathway (rather than the DSBR pathway).[1] The SDSA pathway produces non-crossover recombinants (see Figure).
In meiosis, the process of crossing-over occurs between non-sister chromatids, which results in new allelic combinations in the gametes. In crossing-over, a Spo11 enzyme makes staggered nicks in a pair of sister chromatid strands (in a tetrad organization of prophase). Subsequent enzymes trim back the 5' ends of the strand and a protein complex binds to the 3' single-stranded ends. Rad51 protein is recruited and binds in a protein complex to search for a complementary sequence analogous to double-strand-break repair. The filament searches for the homologous chromosome, strand invasion occurs where the new chromosome forms a D-loop over the bottom sister chromatid, then the ends are annealed. This process can yield double Holliday junctions that when cut in a transversal pattern by endonucleases form 2 heteroduplex strand products.
Heteroduplex DNA is also a source of small RNAs (smRNAs), causing post-transcriptional gene silencing.
References
[edit]- ^ a b Andersen SL, Sekelsky J (December 2010). "Meiotic versus mitotic recombination: two different routes for double-strand break repair: the different functions of meiotic versus mitotic DSB repair are reflected in different pathway usage and different outcomes". BioEssays. 32 (12): 1058–1066. doi:10.1002/bies.201000087. PMC 3090628. PMID 20967781.
- ^ Allers T, Lichten M (July 2001). "Differential timing and control of noncrossover and crossover recombination during meiosis". Cell. 106 (1): 47–57. doi:10.1016/s0092-8674(01)00416-0. PMID 11461701. S2CID 1878863.
